RNA Epigenome Dynamics
Decoding noncoding RNAs
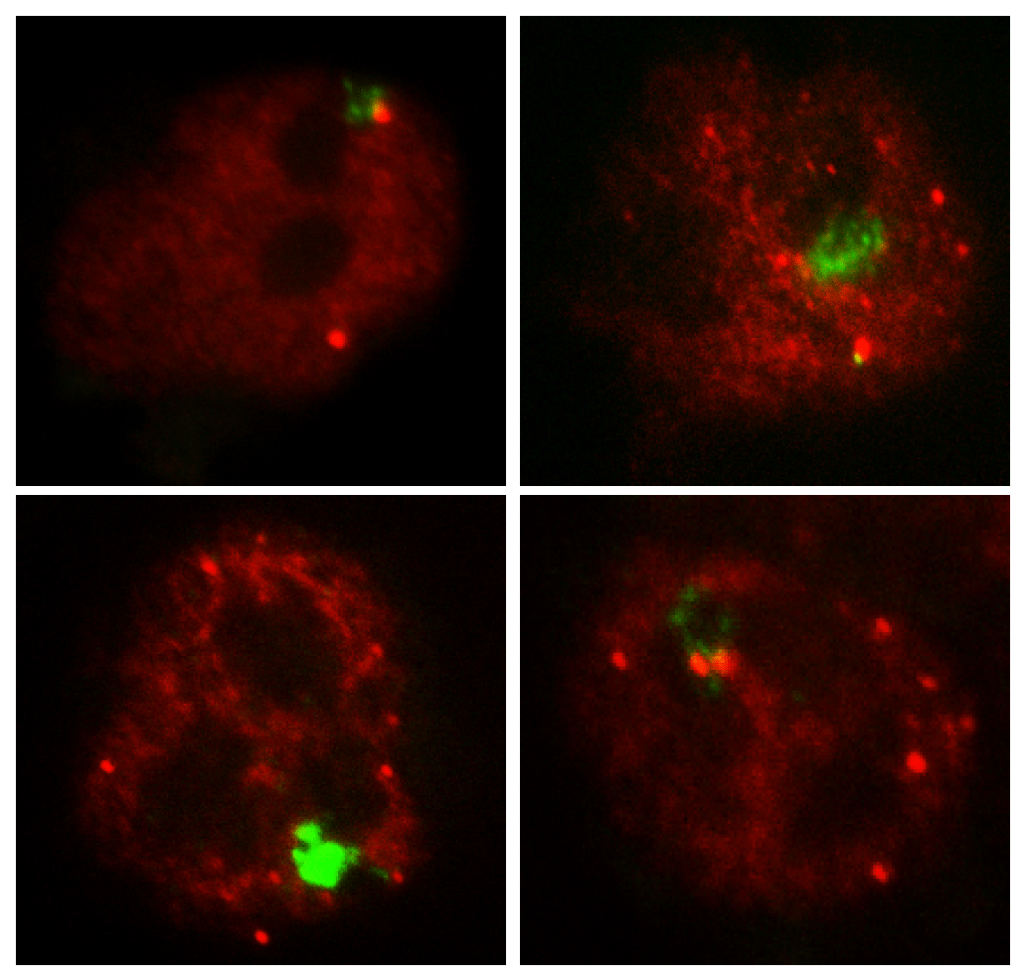
Jpx (red) and Xist (green).
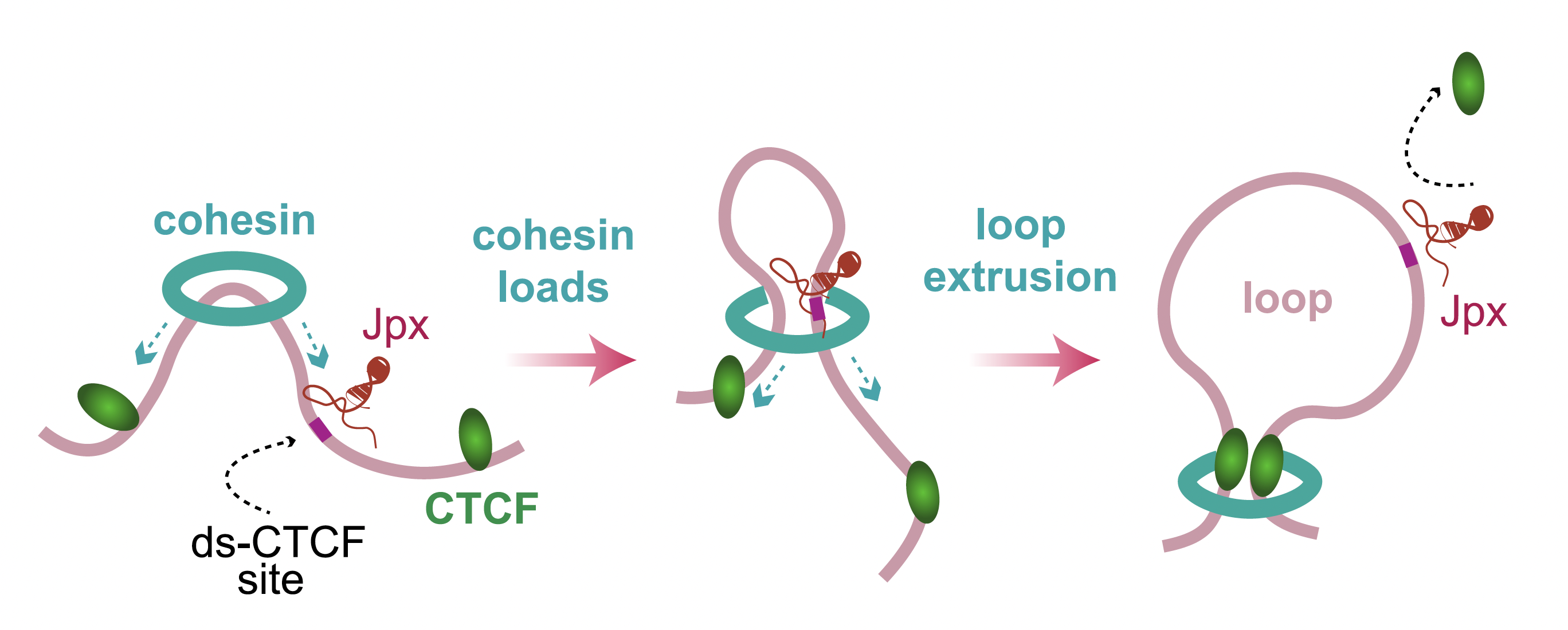
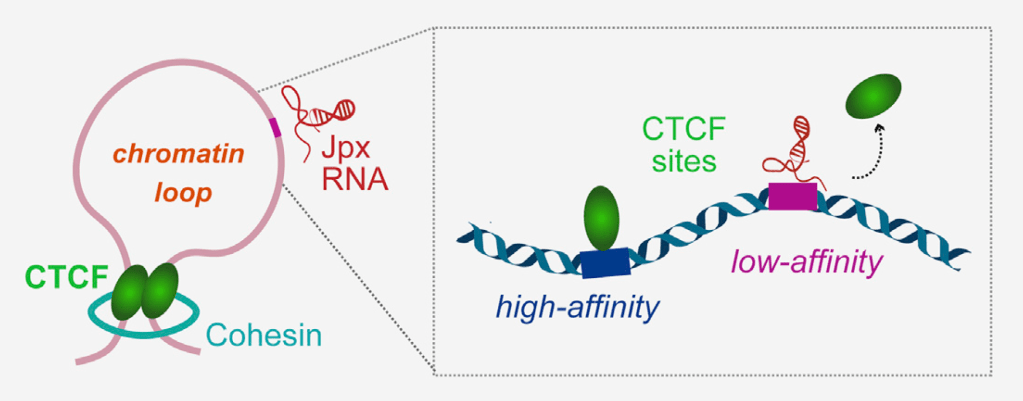
Jpx shapes the 3D genome by regulating anchor site usage
Oh et al. Cell 2021
Unveiling long noncoding RNAs (lncRNAs):
Shaping Epigenomic Architecture
We delve into the fascinating world of long noncoding RNAs (lncRNAs) and their pivotal role in shaping the 3D epigenome dynamics. We aim to unravel the mechanisms of how dysregulated lncRNAs and alterations in chromatin topology lead to pathogenesis. We use embryonic stem cell models, coupled with functional perturbation techniques and multi-omics approach.
Join us to explore lncRNA functions, 3D genome regulation, and profound complexities of gene regulation!
What we do

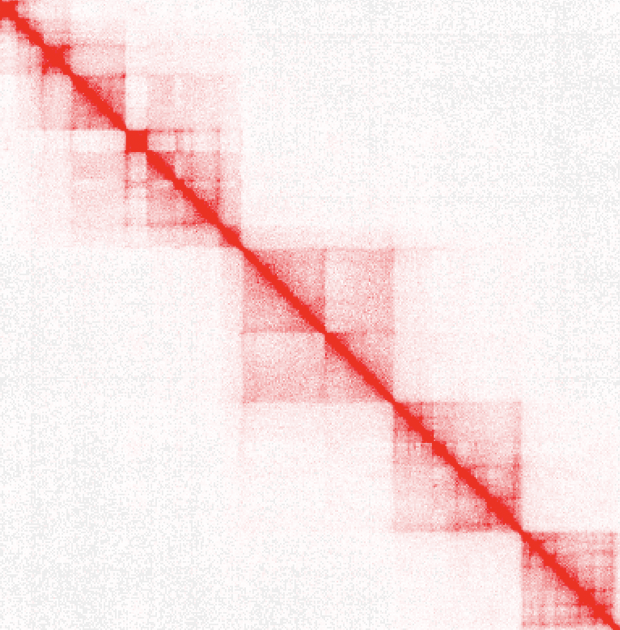
3D genome architecture
We explore the spatial organization of the mammalian genome across various levels. Our aim is to uncover the mechanisms regulating the formation and dynamic alteration of chromosome loops in the 3D genome, with a specific focus on RNA-directed mechanisms.

Long noncoding RNAs
LncRNAs hold the key to understanding the mechanisms that govern highly regulated gene expression in complex organisms. However, their functions largely await exploration. We aim to understand the biology of lncRNAs and the mechanisms of their actions.
Multi-omics
To achieve our goals, we use epigenomics, transcriptomics, proteomics, 3C-based technologies, and bioinformatics.